Finding the information that you need on plant genomes is difficult when published data is spread across a vast amount of journals and the rest is unpublished. So we gathered together 10 of the best plant genome databases to make your job easier.
Brassica.info
 A range of databases exist which manage information relating to Brassica genetics, genomics and related activities. There remains a challenge to extend the scope of nomenclature standards, management of ‘registries’ of widely used resources and information, and development of tools for integrating and navigating between disparate datasets.
A range of databases exist which manage information relating to Brassica genetics, genomics and related activities. There remains a challenge to extend the scope of nomenclature standards, management of ‘registries’ of widely used resources and information, and development of tools for integrating and navigating between disparate datasets.
Ensembl Plants
Ensembl Plants is a genome-centric portal for plants of scientific interest, providing statistics (for example, base pairs and genome size) and information on assembly, annotation, regulation, variation and sequences. Data can be searched by gene identifier or by BLAST sequence search.
Genome Size in Asteraceae Database
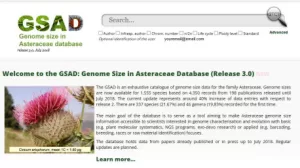
The GSAD is an exhaustive catalogue of genome size data for the family Asteraceae. Genome sizes are now available for 1,555 species based on 4,350 records from 198 publications released until July 2018. The current update represents around 40% increase of data entries with respect to release 2. There are 337 species (21.67%) and 46 genera (19,83%) recorded for the first time.
NCBI
The National Center for Biotechnology Information provides access to genomic information in order to advance science. The genome database currently comprises of 16,326 species and can be sorted by kingdom, group and subgroup.
Plant Garden

The Plant Garden Database brings together information from smaller databases and literature. Information on the organism, genome (for example, chromosome number and genome size), markers and genome specific databases can be accessed.
Phytozome
Phytozome is a plant comparative genomics portal wherein families of related genes representing the modern descendants of ancestral genes are constructed at key phylogenetic nodes. Currently Phytozome provides access to 58 sequenced and annotated green plant genomes.
Plant DNA C-values database
Kew Royal Botanic Garden’s Plant DNA C-values database contains C-values for 8,510 angiosperm, gymnosperm, pteridopghyte, bryophyte and algal species. Users can choose between analysing C-value data across different groups of plants, or searching just part of the database by selecting the specific plant group of interest.
Plant rDNA Database

The Plant rDNA database is an online resource providing information on numbers and positions of ribosomal DNA signals and their structures for 2770 plant species (4948 entries). The data have been obtained from 785 publications on plant molecular cytogenetics using mostly fluorescent in situ hybridisation (FISH) of the 5S and the 18S-5.8S-26S ribosomal RNA genes (rDNA). Where available it also includes information on ploidy level, chromosome number, genome size and life cycle.
Plant GDB Genome Browser

The Plant GDB Genome Browser displays current gene structure models and transcript evidence from spliced alignments of EST and cDNA sequences. If applicable, matched GSS contigs, community annotations, similar proteins, and microarray probes are displayed.
BnaOmics
The BnaOmics database allows you to search genes and transcripts by species, dataset, genome location, name and/or keyword. For gene ID, you can just input a gene ID and begin to search. For genome location, you need to select a dataset and chromosome to search. For keyword, enter any protein name of homologs, GO term, or InterPro term.





